Main data types
One dimensional vectors: double, integer, logical, complex, characters.
integer and double are both numerical
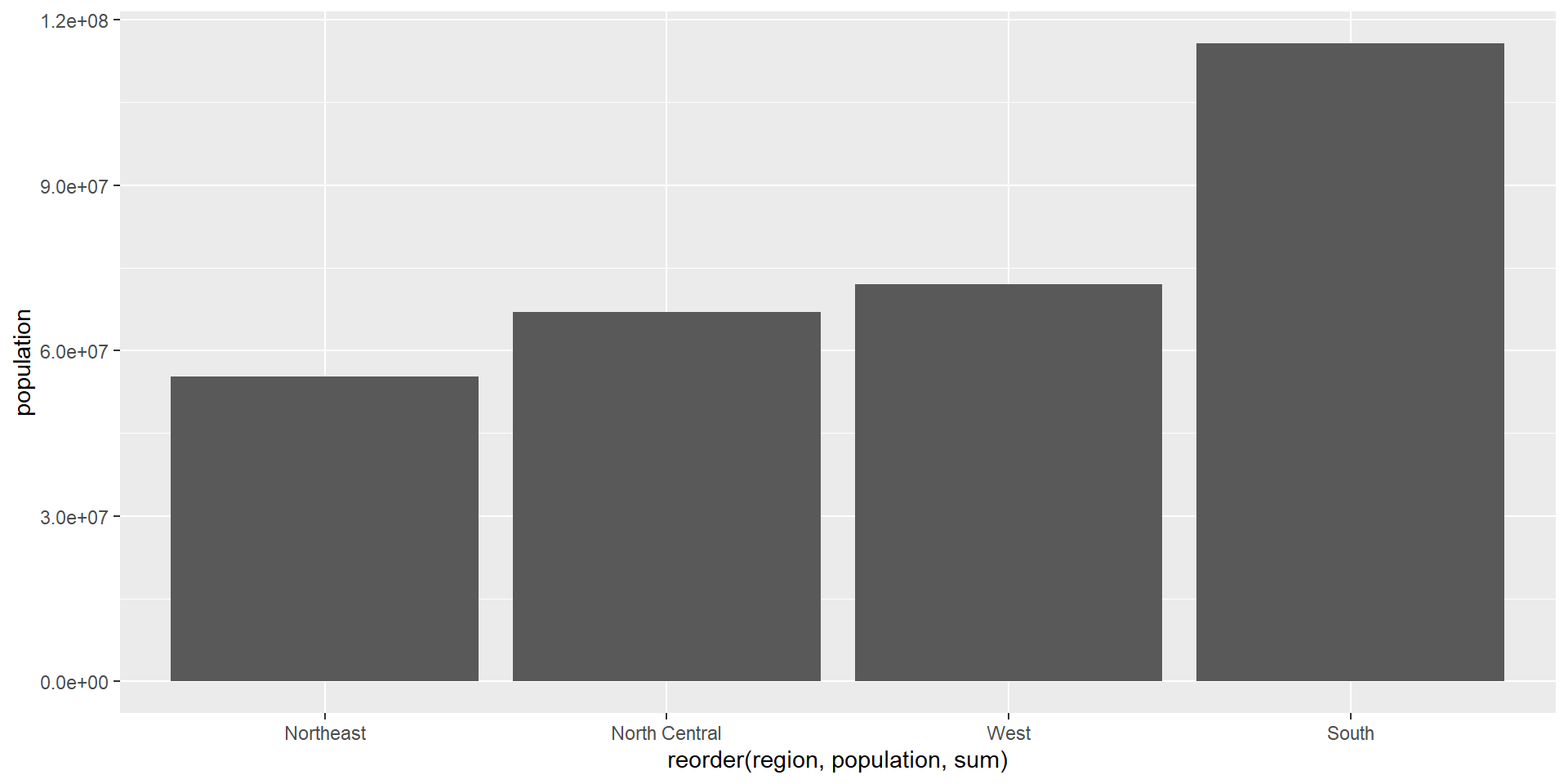
Factors (categorical variables): take finite discrete values
Lists: this includes data frames.
Arrays (matrices): all elements are of the same type
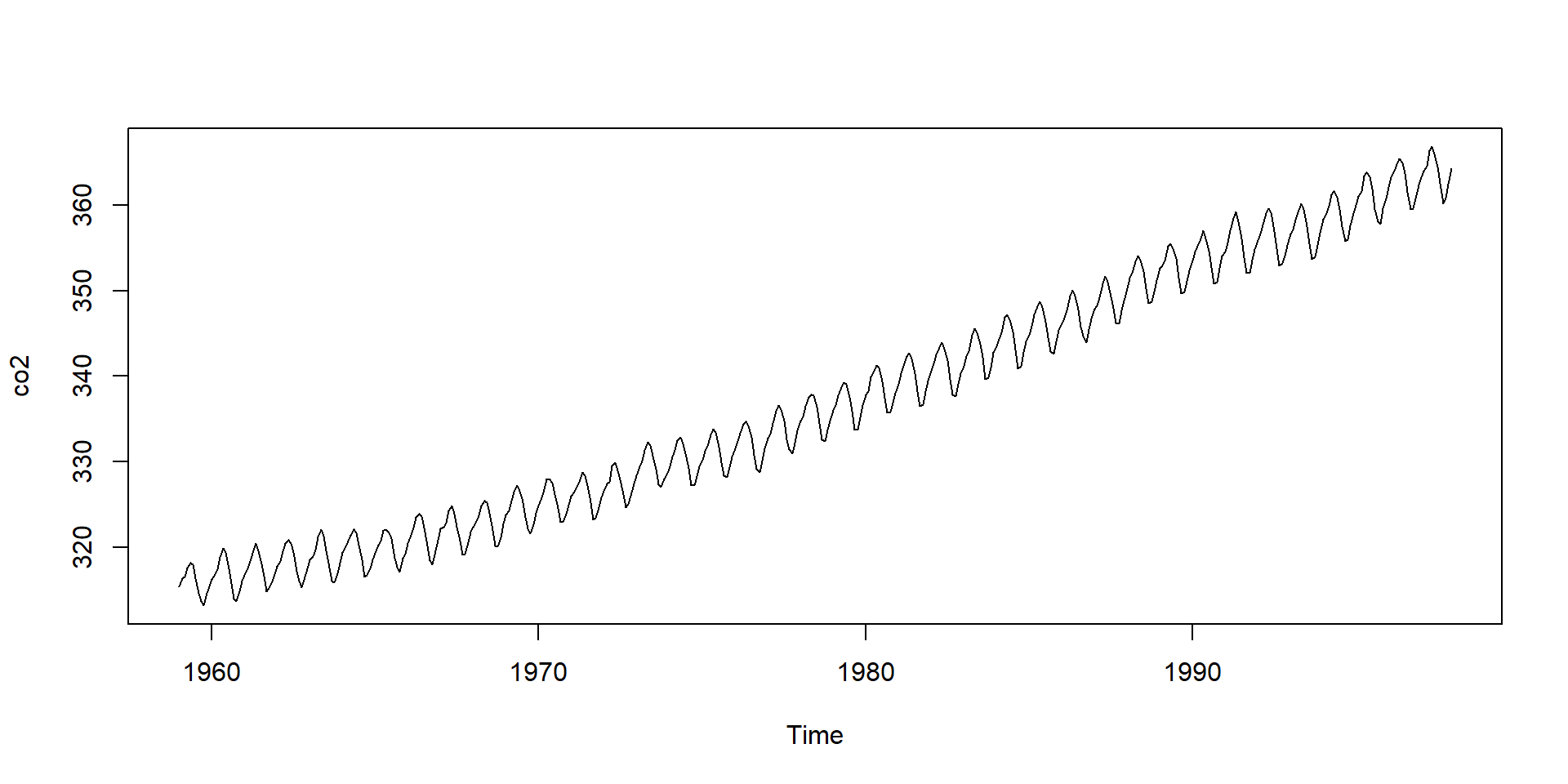
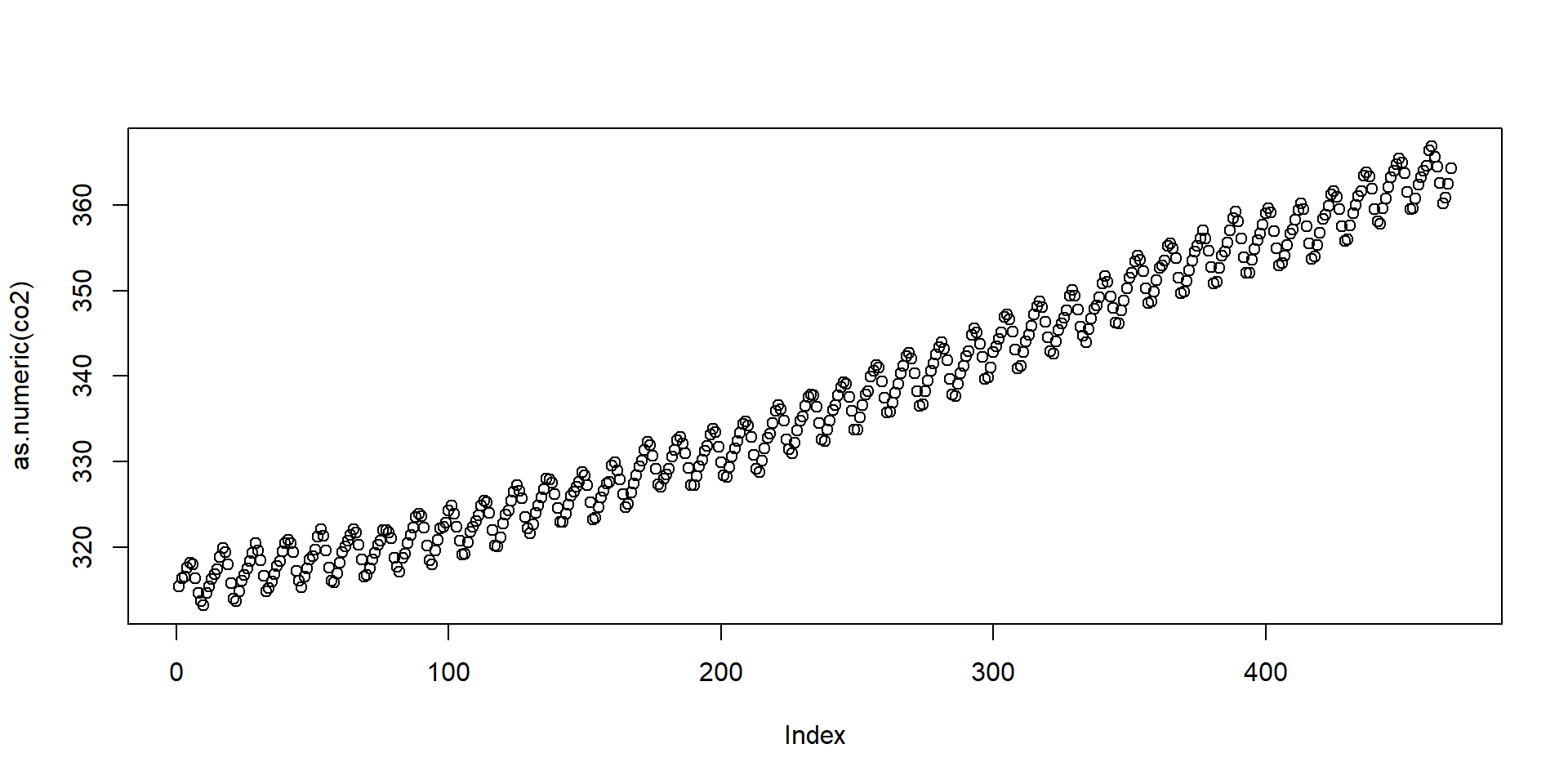
Date and time
tibble (comes from
tidyverse, behaves like a dataframe, print friendly, no automatic type change, avoids silent bugs)S3 objects (default R system, flexible, but can be fragile) vs. S4 objects (srongly typed classes, avoid silent bugs)